UBA protein domain
| UBA | |||||||||
|---|---|---|---|---|---|---|---|---|---|
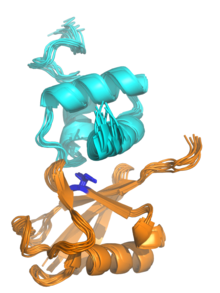 The NMR structure of a UBA domain from the protein ubiquilin-1 (top, cyan) bound to ubiquitin (bottom, orange), illustrating the three-helix bundle structure of the UBA domain. Isoleucine 44, the center of a hydrophobic patch on the ubiquitin surface that interacts with a number of ubiquitin-binding domains, is highlighted in blue. Rendered from PDB: 2JY6.[1] | |||||||||
| Identifiers | |||||||||
| Symbol | UBA | ||||||||
| Pfam | PF00627 | ||||||||
| Pfam clan | CL0214 | ||||||||
| ECOD | 103.1.1 | ||||||||
| InterPro | IPR015940 | ||||||||
| PROSITE | PDOC50030 | ||||||||
| SCOP2 | 1efu / SCOPe / SUPFAM | ||||||||
| CDD | cd00194 | ||||||||
| |||||||||
Ubiquitin-associated (UBA) domains are protein domains that non-covalently interact with ubiquitin through protein-protein interactions. Ubiquitin is a small protein that is covalently linked to other proteins as part of intracellular signaling pathways, often as a signal for protein degradation. UBA domains are among the most common ubiquitin-binding domains.[2][3]
Function
[edit]Proteins containing UBA domains are involved in a variety of additional cell processes, such as nucleotide excision repair (NER), spindle pole body duplication, and cell growth.[4]
Protein degradation via the ubiquitin proteasome system (UPS) allows the cell to selectively negatively regulate intracellular proteins. Protein degradation helps to maintain protein quality control, signalling, and cell cycle progression.[5][6] UBA has been proposed to limit ubiquitin chain elongation and to target polyubiquitinated proteins to the 26S proteasome for degradation.[7] They have been identified in modular proteins involved in protein trafficking, DNA repair, proteasomal degradation, and cell cycle regulation.
Structure
[edit]UBA domains have a common sequence motif of approximately 45 amino acid residues.[8] They fold into three-helix bundle structures.[2]
Examples
[edit]
The human homologue of yeast Rad23A is one example of a nucleotide excision-repair protein that contains both an internal and a C-terminal UBA domain. The solution structure of human Rad23A UBA(2) showed that the domain forms a compact three-helix bundle.[10]
Comparison of the structures of UBA(1) and UBA(2) reveals that both form very similar folds and have a conserved large hydrophobic surface patch which may be a common protein-interacting surface present in diverse UBA domains. Evidence that ubiquitin binds to UBA domains leads to the prediction that the hydrophobic surface patch of UBA domains interacts with the hydrophobic surface on the five-stranded beta-sheet of ubiquitin.[11]
This domain is similar in sequence to the N-terminal domain of translation elongation factor EF1B (or EF-Ts) from bacteria, mitochondria and chloroplasts.[9]
References
[edit]- ^ Zhang D, Raasi S, Fushman D (March 2008). "Affinity makes the difference: nonselective interaction of the UBA domain of Ubiquilin-1 with monomeric ubiquitin and polyubiquitin chains". Journal of Molecular Biology. 377 (1): 162–80. doi:10.1016/j.jmb.2007.12.029. PMC 2323583. PMID 18241885.
- ^ a b Dikic I, Wakatsuki S, Walters KJ (October 2009). "Ubiquitin-binding domains - from structures to functions". Nature Reviews. Molecular Cell Biology. 10 (10): 659–71. doi:10.1038/nrm2767. PMC 7359374. PMID 19773779.
- ^ Husnjak K, Dikic I (7 July 2012). "Ubiquitin-binding proteins: decoders of ubiquitin-mediated cellular functions". Annual Review of Biochemistry. 81 (1): 291–322. doi:10.1146/annurev-biochem-051810-094654. PMID 22482907.
- ^ Su V, Lau AF (September 2009). "Ubiquitin-like and ubiquitin-associated domain proteins: significance in proteasomal degradation". Cellular and Molecular Life Sciences. 66 (17): 2819–33. doi:10.1007/s00018-009-0048-9. PMC 2725189. PMID 19468686.
- ^ Gomez TA, Kolawa N, Gee M, Sweredoski MJ, Deshaies RJ (May 2011). "Identification of a functional docking site in the Rpn1 LRR domain for the UBA-UBL domain protein Ddi1". BMC Biology. 9: 33. doi:10.1186/1741-7007-9-33. PMC 3126750. PMID 21627799.
- ^ Tse MK, Hui SK, Yang Y, Yin ST, Hu HY, Zou B, et al. (2011). "Structural analysis of the UBA domain of X-linked inhibitor of apoptosis protein reveals different surfaces for ubiquitin-binding and self-association". PLOS ONE. 6 (12): e28511. Bibcode:2011PLoSO...628511T. doi:10.1371/journal.pone.0028511. PMC 3240630. PMID 22194841.
- ^ Li J, Chu H, Zhang Y, Mou T, Wu C, Zhang Q, Xu J (2012). "The rice HGW gene encodes a ubiquitin-associated (UBA) domain protein that regulates heading date and grain weight". PLOS ONE. 7 (3): e34231. Bibcode:2012PLoSO...734231L. doi:10.1371/journal.pone.0034231. PMC 3311617. PMID 22457828.
- ^ Hofmann K, Bucher P (May 1996). "The UBA domain: a sequence motif present in multiple enzyme classes of the ubiquitination pathway". Trends in Biochemical Sciences. 21 (5): 172–3. doi:10.1016/S0968-0004(96)30015-7. PMID 8871400.
- ^ a b Kawashima, Takemasa; Berthet-Colominas, Carmen; Wulff, Michael; Cusack, Stephen; Leberman, Reuben (8 February 1996). "The structure of the Escherichia coli EF-Tu· EF-Ts complex at 2.5 Å resolution". Nature. 379 (6565): 511–518. Bibcode:1996Natur.379..511K. doi:10.1038/379511a0. PMID 8596629. S2CID 4273375.
- ^ Dieckmann T, Withers-Ward ES, Jarosinski MA, Liu CF, Chen IS, Feigon J (December 1998). "Structure of a human DNA repair protein UBA domain that interacts with HIV-1 Vpr". Nature Structural Biology. 5 (12): 1042–7. doi:10.1038/4220. PMID 9846873. S2CID 30478711.
- ^ Mueller TD, Feigon J (June 2002). "Solution structures of UBA domains reveal a conserved hydrophobic surface for protein-protein interactions". Journal of Molecular Biology. 319 (5): 1243–55. doi:10.1016/S0022-2836(02)00302-9. PMID 12079361.
